Background: Guselkumab (GUS), an interleukin-23 p19-subunit monoclonal antibody, demonstrated efficacy compared with placebo (PBO) in reducing signs and symptoms of psoriatic arthritis (PsA) in the phase 3 DISCOVER-1 & 2 studies. 1,2
Objectives: To evaluate gene expression in the blood of PsA patients (pts) in the DISCOVER-1 & -2 studies and the impact of GUS on the expression of these genes.
Methods: Pts were treated with GUS 100 mg every 4 weeks (Q4W); GUS 100 mg at W0, W4, then Q8W; or matching PBO. Whole transcriptome profiling by RNA-sequencing was performed using the Novaseq platform on blood samples obtained from a subset of 673 pts with PsA at baseline across the 2 DISCOVER studies, as well as from 21 demographically (age, sex, and ethnicity) matched healthy controls procured independently of the clinical program. A subgroup (N=227) also had serial blood samples (W0/W4/W24) evaluated; the subgroup pts were selected based on having baseline characteristics (demographics, disease activity, medication use) representative of the overall cross-study PsA population. Significance of differentially expressed genes (DEGs) between PsA and healthy controls was defined by a false discovery rate (FDR) <0.05 based on a log-linear model using edgeR. Top genes were defined by significance and |logFC| >1. For cell type analysis, genes that changed with GUS treatment were tested for enrichment using Cibersort. Gene enrichment scores were calculated using Gene Set Variation Analysis (GSVA).
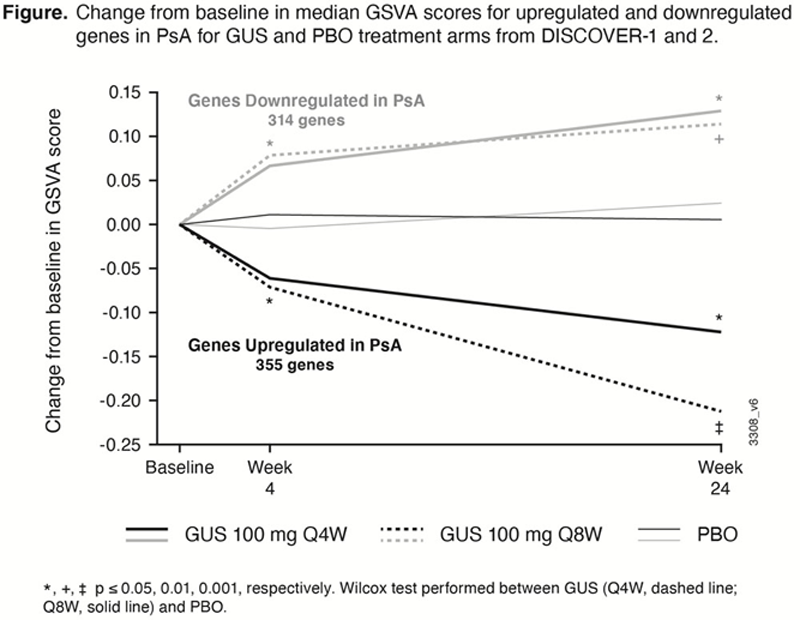
Results: To define disease genes, we compared genes at baseline in pts with active PsA vs. healthy control whole blood transcriptomes and detected 355 upregulated and 314 downregulated (top genes shown in
Conclusion: Using whole transcriptome profiling, we detected DEGs in blood samples obtained from PsA pts vs. healthy controls, suggesting a dysregulation of immune cell profiles in PsA. The majority of these disease-associated genes were modulated by GUS, with directionality toward a normalization of whole blood transcriptomic signatures.
REFERENCES:
[1]Deodhar A et al. Lancet. 2020;395:1115.
[2]Mease P et al. Lancet. 2020;395:1126.
Top DEGs derived from PsA vs. healthy whole blood transcriptomes.
| Upregulated in PsA | Downregulated in PsA | ||||||
| Gene | logFC | logCPM | FDR | Gene | logFC | logCPM | FDR |
| ADGRG7 | 5.92 | -0.90 | 0.02101 | AK8 | -1.36 | -1.06 | 1.61E-07 |
| ADAMTS2 | 4.06 | 0.82 | 0.006466 | FTCD | -1.48 | -1.74 | 1.67E-05 |
| PGF | 3.21 | -0.68 | 0.006466 | GPR15 | -1.54 | 1.81 | 1.67E-05 |
| PCSK9 | 3.21 | -2.96 | 0.023872 | CHRM3 | -1.54 | -2.62 | 9.6E-08 |
| OLAH | 2.76 | 0.75 | 0.004539 | RFPL4AL1 | -1.69 | -3.34 | 0.009738 |
| MAOA | 2.55 | -0.26 | 0.005463 | SPACA3 | -1.85 | -3.23 | 0.000216 |
| SLC2A14 | 2.30 | 0.59 | 0.022594 | VANGL2 | -1.95 | -1.79 | 9.6E-08 |
| MMP1 | 2.25 | -1.16 | 0.004745 | RFPL4A | -2.04 | -1.28 | 0.004539 |
| DAAM2 | 2.12 | 4.31 | 0.024628 | GLYATL2 | -2.77 | -2.78 | 1.93E-15 |
| BCAR1 | -3.13 | -2.58 | 6.24E-26 | ||||
Bold indicates positive change. CPM = counts per million.
Disclosure of Interests: Stefan Siebert Consultant of: AbbVie, Janssen, Novartis, UCB, Grant/research support from: AbbVie, Amgen (previously Celgene), Bristol Myers Squibb, Boehringer Ingelheim, GSK, Janssen, Novartis, UCB, Kristen Sweet Shareholder of: Johnson & Johnson, Employee of: Janssen Research & Development LLC, Christopher T. Ritchlin Consultant of: AbbVie, Amgen, Gilead, Janssen, Eli Lilly, Novartis, Pfizer, and UCB, Grant/research support from: AbbVie, Amgen, and UCB, Elizabeth C Hsia Shareholder of: Johnson & Johnson, Employee of: Janssen Research & Development LLC, Alexa Kollmeier Shareholder of: Johnson & Johnson, Employee of: Janssen Research & Development LLC, Xie L Xu Shareholder of: Johnson & Johnson, Employee of: Janssen Research & Development LLC, Qingxuan Song Shareholder of: Johnson & Johnson, Employee of: Janssen Research & Development LLC, Michelle Miron Shareholder of: Johnson & Johnson, Employee of: Janssen Research & Development LLC