

Background: Metabolic pathways are important regulators of immune differentiation and activation in kidneys. Kidneys directly impact systemic metabolism, circulating metabolite levels, and express intrinsic metabolic activity. The integration of renal metabolomic and transcriptomic profiles may unravel unique gene-metabolite pairs of biological significance in lupus nephritis (LN).
Objectives: To decipher gene-metabolite signatures at both pre-nephritic and nephritic stages of lupus.
Methods: Kidneys were isolated and snap-frozen after perfusion from female NZB/NZW-F1 lupus mice at the pre-nephritic (3-month-old) and nephritic (6-month-old exhibiting ≥100 ng/dL of urine protein) stage of lupus (n=6/group). Age-matched female C57BL/6 mice were used as healthy controls. Sample extracts were used for RNA sequencing and 1 H-NMR spectroscopy metabolic profiling. DESeq2 was used to identify differentially expressed genes. Univariate analysis was used to reveal metabolic differences characteristic for nephritis.
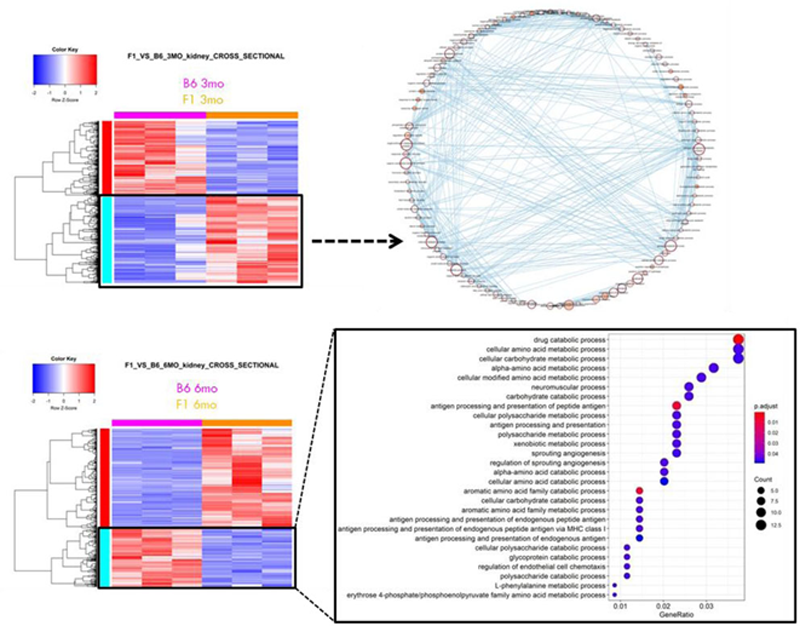
Results: Comparative transcriptomic analyses uncovered multiple transcripts related to metabolic pathways: In pre-nephritic kidneys, lipid metabolism, cellular respiration, TCA cycle, amino acid metabolism processes were overrepresented in the upregulated genes while in nephritic kidneys, amino acid metabolism processes were overrepresented among the downregulated genes (
Conclusion: The combined transcriptomics and metabolomics analysis revealed metabolic derangements in lupus-affected kidneys both during subclinical and overt LN. Deregulated tissue-levels of taurine and myo-inositol at the subclinical stage of the disease suggest aberrant renal biochemistry preceding the development of overt LN that may directly impact systemic metabolism and circulating metabolite levels.
Pathways linked to cell metabolism were overrepresented among 3-month upregulated and 6-month lupus mice (F1) downregulated DEGS ( differentially expressed genes ) compared to controls (C57BL/6).
Acknowledgements: This project has received funding from the European Research Council (ERC) under the European Union’s Horizon 2020 research and innovation programme (grant agreement No 742390).
Disclosure of Interests: None declared