

Background: Anti-SSA/Ro autoantibodies are among the most frequently detected extractable nuclear antigen autoantibodies and have mainly been associated with primary Sjögren’s syndrome (pSS), systemic lupus erythematosus (SLE) and undifferentiated connective tissue disease (UCTD).
Objectives: Is there a common signature to all patients expressing anti-Ro60 autoantibodies regardless of their disease phenotype?
Methods: Using high-throughput multi-omics data collected within the cross-sectional cohort from the PRECISESADS IMI project [1] (genetic, epigenomic, transcriptomic, combined with flow cytometric data, multiplexed cytokines, classical serology and clinical data), we assessed by machine learning the integrated molecular profiling of 520 anti-Ro60-positive (anti-Ro60 + ) compared to 511 anti-Ro60-negative (anti-Ro60 - ) patients with pSS, SLE and UCTD, and 279 healthy controls (HCs).
Results: The selected features for RNA-Seq, DNA methylation and GWAS data allowed a clear separation between anti-Ro60
+
and anti-Ro60
-
patients. These results demonstrate that the different features selected by machine learning from the anti-Ro60
+
patients constitute specific signatures when compared to anti-Ro60
-
patients and HCs. Remarkably, the gene transcript
z
-score of three genes (
ATP10A
,
MX1
and
PARP14
), presenting an overexpression associated with a hypomethylation and genetic variation, and independently identified by the Boruta algorithm, was clearly higher in anti-Ro60
+
patients compared to anti-Ro60
-
patients in all the diseases (
Three genes common to RNA-Seq, DNA methylation and GWAS analysis characterize anti-Ro60 + patients. (A) ATP10 / MX1 / PARP14 z -score analyses were performed for 731 patients and 254 HCs according to anti-Ro60 expression. (B) ATP10 / MX1 / PARP14 z -score analyses were performed for 286 pSS, 351 SLE and 94 UCTD patients and 254 HCs. Two-tailed pairwise Wilcoxon-rank sum test results are shown. Plots show median, with error bars indicating ± interquartile range. (pSS: primary Sjögren’s syndrome, SLE: systemic lupus erythematosus, UCTD: undifferentiated connective tissue disease, HCs: healthy controls).
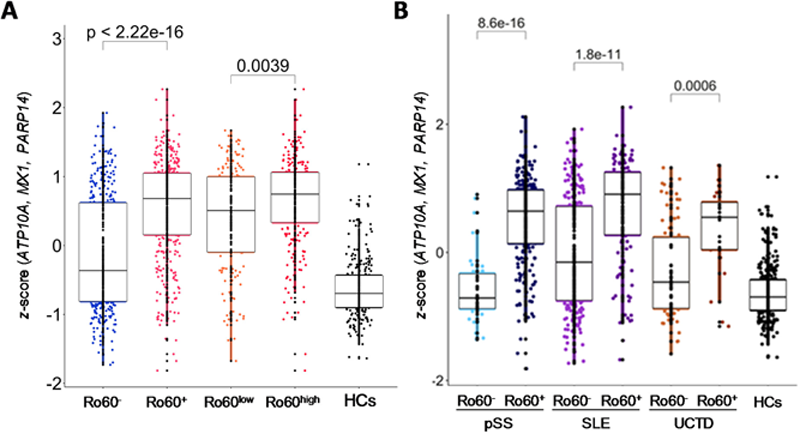
Conclusion: Anti-Ro60 + patients present a specific inflammatory signature regardless of their disease suggesting that a dual approach targeting both Ro-associated RNAs and anti-Ro60 autoantibodies should be considered.
REFERENCES:
[1]Barturen G, Babaei S, Català-Moll F, Martínez-Bueno M, Makowska Z, Martorell-Marugán J, et al. Integrative Analysis Reveals a Molecular Stratification of Systemic Autoimmune Diseases. Arthritis Rheumatol Hoboken NJ. 2021 Jun;73(6):1073–85
Funding: The research leading to these results has received support from the Innovative Medicines Initiative Joint Undertaking under the Grant Agreement Number 115565 (PRECISESADS project), resources of which are composed of financial contribution from the European Union’s Seventh Framework Program (FP7/2007–2013) and EFPIA companies’ in-kind contribution.
Acknowledgements: The study has been conducted thank to the contribution of the PRECISESADS clinical consortium and the PRECISESADS flow cytometry consortium.
Disclosure of Interests: None declared.