

Background: Rheumatoid arthritis (RA) is a complex disease, caused by a combination of genetic, epigenetic and environmental factors common to other related autoimmune diseases including Multiple Sclerosis (MS) and Systemic Lupus Erythematosus (SLE) [1]. Using state of the art Bioinformatics tools we are able to formulate an ensemble of associated components (genomic grammar) for each disease and distinguish important differences and common aspects in a specific group of disease such as ensembles of autoimmune diseases [2].
Objectives: To create, collect and evaluate the most credible and unique gene variants, epigenetic variants and single nucleotide polymorphisms (SNPs) causing the basis of an immune disease (the genomic grammar of the disease), which could potentially assist in the process of the RA disease prevention, diagnosis and treatment [3].
Methods: RA related publications from the PubMed have been analyzed using data mining and semantic techniques towards extracting the candidate causative SNPs. The extracted knowledge has been filtered, evaluated, annotated, and classified in a structured database which also includes GWAS information regarding SNPs. Additional clinical, genomic, structural, functional and biological information was also extracted from biological databases including dbSNP, LitVar, ClinVar and OMIM and cross-correlated with other available autoimmune disease related SNP databases, including the Demetra application, Epione application and Panacea application databases [3, 4].
Results: A holistic genetic map of the studied autoimmune diseases with more than 2000 related SNPs has been estimated and specific sub-clusters with crucial nodes have been identified across the RA, SLE and MS diseases. Based on these results, the three studied autoimmune diseases share a 10% common SNPs genetic background (
Common Related Genes based on the analyzed SNP targets in the studied disease.
| A/A | Gene / Region | A/A | Gene / Region |
|---|---|---|---|
| 1. | ADAM33 | 2. | LOC285626 |
| 3. | ADIPOQ | 4. | MIR3142HG |
| 5. | CD40 | 6. | MIR499A |
| 7. | CIITA | 8. | MTHFR |
| 9. | CTLA4 | 10. | MT-ND5 |
| 11. | FCRL3 | 12. | NCF1 |
| 13. | HLA-DPB1 | 14. | NLRP1 |
| 15. | HLA-DRA | 16. | NOS3 |
| 17. | HLA-G | 18. | NR3C1 |
| 19. | IL17A | 20. | PADI4 |
| 21. | IL1RN | 22. | PDCD1 |
| 23. | IL2 | 24. | PON1 |
| 25. | IL23R | 26. | STAT4 |
| 27. | IL6 | 28. | TGFB1 |
| 29. | IL7R | 30. | TLR9 |
| 31. | IRAK1 | 32. | TNF |
| 33. | VDR | 34. | TNFRSF1A |
| 35. | IRF5 | 36. | TYK2 |
| 37. | KIF5A | 38. | UCP2 |
| 39. | LEP |
Three class Venn diagram of the genomic grammar between RA, MS and SLE.
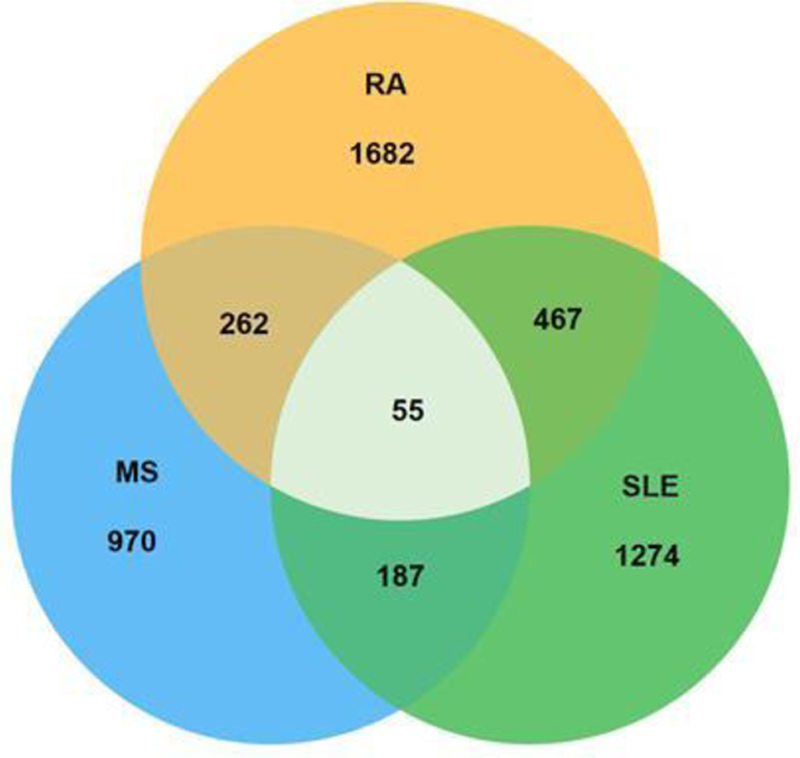
Conclusion: The identification of the optimal genomic grammar in RA will help towards understanding the nature of the disease. Specific genetic targets via determined SNPs could act as biomarkers that aid in forming the right diagnosis [6].
REFERENCES:
[1]Acosta-Herrera et al, J Clin Med 2019;8:826
[2]Chatzikyriakidou et al, Semin Arthritis Rheum 2013;43:29
[3]Papageorgiou et al, Int J Mol Med 2021;47:115
[4]Papageorgiou et al, Int J Mol Med 2022;49:8
[5]Wang, Y et al, Ann Rheum Dis 2021Epub ahead of print:doi:10.1136/annrheumdis-2021-220066
[6]Kurko et al, Clin Rev Allergy Immunol 2013;45:170
Disclosure of Interests: None declared