

Background: Systemic lupus erythematosus (SLE) is a chronic relapsing autoimmune disorder that is challenging to diagnose due to its heterogeneous presentation of a combination of non-specific clinical manifestations and laboratory findings. This heterogeneity and lack of granularity is a major cause of treatment failure. With the advent of precision medicine, there is a need for accurate methods to identify cases of SLE in large clinical databases, and to recognize constellations of pathological characteristics (or fingerprints ) that allow a better understanding of the disease. Computational phenotype discovery aims to solve this problem by using unbiased, data-driven machine learning methods to disentangle the fingerprints of disease that hide within the noisy, sparse and incomplete electronic health record data.
Objectives: To generate data-driven phenotypic fingerprints of SLE and validate them by assessing their ability to distinguish records of patients with SLE vs. ‘near miss’ patients without the disease.
Methods: Records of 716 patients that published, expert-curated algorithms [1] indicated as likely SLE patients were reviewed by 3 clinicians and labeled as positive (490 records), negative (261), or indeterminate (55). Those labeled positive by an algorithm but negative by clinician chart review were considered ‘near misses’ and should be among the more difficult cases to classify. We trained an ElasticNet (regularized logistic regression) model to distinguish the true positive records from the near misses and evaluated it under 10x cross validation.
Inputs to the predictive model were the projections of each patient record into the space of 2000 latent variables. These features had been inferred separately by an unsupervised machine-learning algorithm from a much larger dataset of 646,716 randomly sampled temporal cross sections of 63,775 longitudinal records of patients with an antinuclear antibody test. Each of the 2000 variables constitutes the phenotypic fingerprint of an unobserved, independent potential disease mechanism. Formally, they represent linear combinations of demographic data, laboratory test results, billing code intensities, and medication exposures [2].
Results: Our predictive model achieved an area under the Receiver Operating Characteristic Curve of 0.90, 95% CI: [0.879, 0.922], an area under the Precision-Recall Curve of 0.94, 95% CI: [0.926, 0.954], and an Integrated Calibration Index of 0.074, 95% CI: [0.067, 0.080].
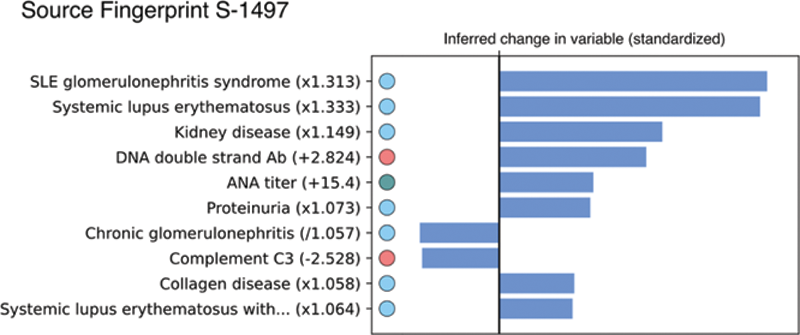
The model selected 61 of the 2000 potential latent mechanisms to distinguish positive SLE records from near-misses. All of the phenotypes selected to be predictive exhibit high face-validity from a clinical interpretation perspective. They include recognizable patterns of variables representing different clinical features of SLE (
Detail of the estimated changes in the clinical features representing one of the 61 potential fingerprints found to distinguish records of patients with SLE vs. near misses.
Conclusion: We found unbiased, data-driven fingerprints of potential latent disease mechanisms that were highly informative in differentiating records of patients with SLE from near-miss presentations, with performance surpassing existing expert-curated algorithms. Our methods found dozens of variable constellations that may lead to a more precise understanding of this complex and heterogeneous disease. That these results were obtained from cases that should be the most difficult for an algorithm to distinguish (positive vs. near-miss) suggests that our model’s accuracy is a lower bound on what would be expected from a more balanced cohort.
REFERENCES:
[1]Barnado et al. Developing Electronic Health Record Algorithms That Accurately Identify Patients With Systemic Lupus Erythematosus. Arthritis care & research, 2017
[2]Thomas A. Lasko and Diego A. Mesa. Computational Phenotype Discovery via Probabilistic Independence. Proc KDD Appl Data Sci for Healthcare (DSHealth), 2019.
Acknowledgements: MBM would like to acknowledge the Fulbright Spanish Comission for funding his graduate studies and the research that made this work possible. VKK would like to acknowledge NIAMS/NIH R01 AR076516 and Lupus Research Alliance BMS Accelerator Award for their financial support.
Disclosure of Interests: None declared