

Background: Skin involvement is common in systemic lupus erythematosus (SLE), with up to 70% of patients developing skin lesions at some point during the course of their disease. Moreover, cutaneous lesions are the first sign of systemic disease in 20% of SLE patients [1]. Skin requires a regulated and undisrupted miRNA profile for a correct development and function. Studies show that miRNAs play a pathogenic role in autoimmune skin diseases such as Psoriasis [2]. Understanding of the role of deregulated miRNAs in CLE may enhance our knowledge about CLE pathogenesis which is currently not understood.
Objectives: Identify differentially expressed miRNAs in skin lesions to determine their role in CLE pathogenesis.
Methods: Paired skin biopsies from CLE lesional (n=20) and non-lesional skin (n=20) were obtained. MiRNA microarray screening was performed using TaqMan MicroRNA Array. Validation was performed in a new cohort of FFPE skin samples by qRT-PCR (n=20) . In situ hybridization was performed to identify the skin cell type involved in selected miRNAs. Next, in vitro experiments using primary skin and immune cells were performed in order to discover their role in CLE pathogenesis. A microarray screening and luciferase assay were performed to find miR-885-5p altered gene expression, molecular pathways and target genes.
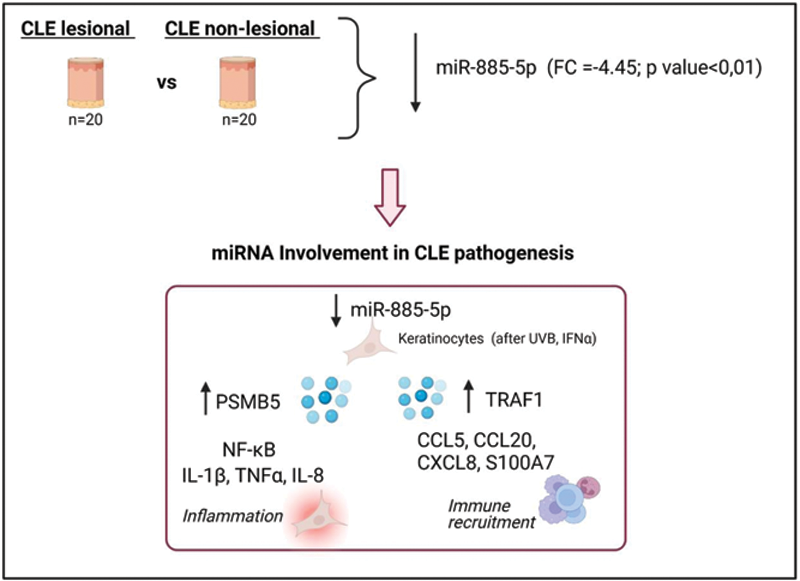
Results: MiR-885-5p was found downregulated in CLE lesional skin (-4.45 fold, pvalue<0,01, respectively). Results were confirmed in a validation cohort. In situ hybridization revealed miR-885-5p was located in the epidermal keratinocytes in CLE non-lesional skin. Healthy keratinocytes presented an increased miR-885-5p expression compared with other relevant cell types such as fibroblasts (p<0.001) or PBMCs (p<0.001). Moreover stimulation with IFNα and UVB promoted a marked long-lasting downregulation of miR-885-5p (IFNα, 24h -67 fold (p<0.001); UVB, 24h -70 fold (p<0.001)).
Gene microarray of anti-miR-885-5p keratinocytes showed that the unique gene that was upregulated in all anti-miR-885-5p conditions was PSMB5. In vitro experiments demonstrated that PSMB5 is a direct target of miR-885-5p (74% of reduction p<0.001) and it is involved in epidermal inflammation by enhancing NFKB and inflammatory cytokines IL-1β, TNFα and CXCL8. In addition, we observed immune recruitment in UVB stimulated anti-miR885-5p keratinocytes and we found that this immune migration is mediated via miR-885-5p direct binding of TRAF1 and subsequent chemokine CCL5, CCL20, CXCL8 and S100A7 secretion.
Conclusion: Our findings demonstrate that CLE lesions present a downregulation of miR-885-5p that is strongly implicated in CLE pathogenesis. Moreover, miRNA applicability as therapeutics makes the discovered miRNAs promising candidates for novel therapy in CLE, as there is a need to develop new therapies for refractory cases and to avoid current treatment significant side effects.
REFERENCES:
[1]Okon LG, Werth VP. Cutaneous lupus erythematosus: diagnosis and treatment. Best Pract Res Clin Rheumatol. 2013; 27(3):391-404.
[2]Sonkoly E et al. MicroRNAs: novel regulators involved in the pathogenesis of psoriasis? PLoS One. 2007. 11;2(7):e610.
Disclosure of Interests: None declared