

Background: Modic type 1 changes (MC1) are changes of vertebral bone marrow signal intensity found predominantly in patients with degenerated intervertebral discs and damaged cartilage endplates. Discs adjacent to MC1 (MC1 discs) degenerate faster and release more pro-inflammatory and pro-fibrotic factors compared to other degenerated discs. Yet, it remains unknown if the degenerative process of MC1 discs differ from non-MC discs.
Objectives: The aim of this study was to identify MC1-specific mechanisms of disc degeneration. We hypothesized that the MC1 discs contain more cleaved extracellular matrix (ECM) fragments and that they are generated through a MC1-specific mechanism.
Methods: Twenty-three degenerated non-MC discs and 30 MC1 discs from patients undergoing lumbar spinal fusion were analyzed. First, shotgun mass spectrometry (MS) was used for proteome analysis, specifically to identify overexpressed proteases. Second, the degradome (all cleaved proteins) was determined with N-terminal amine isotopic labeling of substrates (N-TAILS) MS. The proteins with the greatest numbe r of de-novo fragments were identified. Third, the full-length recombinant proteins were digested with high temperature requirement A serine peptidase 1 (HTRA1), which has been found to be overexpressed in MC1 discs, with the aim of reproducing the protein fragments identified in the human samples. Cleavage was confirmed with SDS-PAGE followed by Coomassie Blue staining. To test the similarity between the in-vitro generated protein fragments and the ones found in human disc samples, the in-vitro generated protein fragments were analysed with N-TAILS. The identified cleavage sites were then compared with those identified in the human samples.
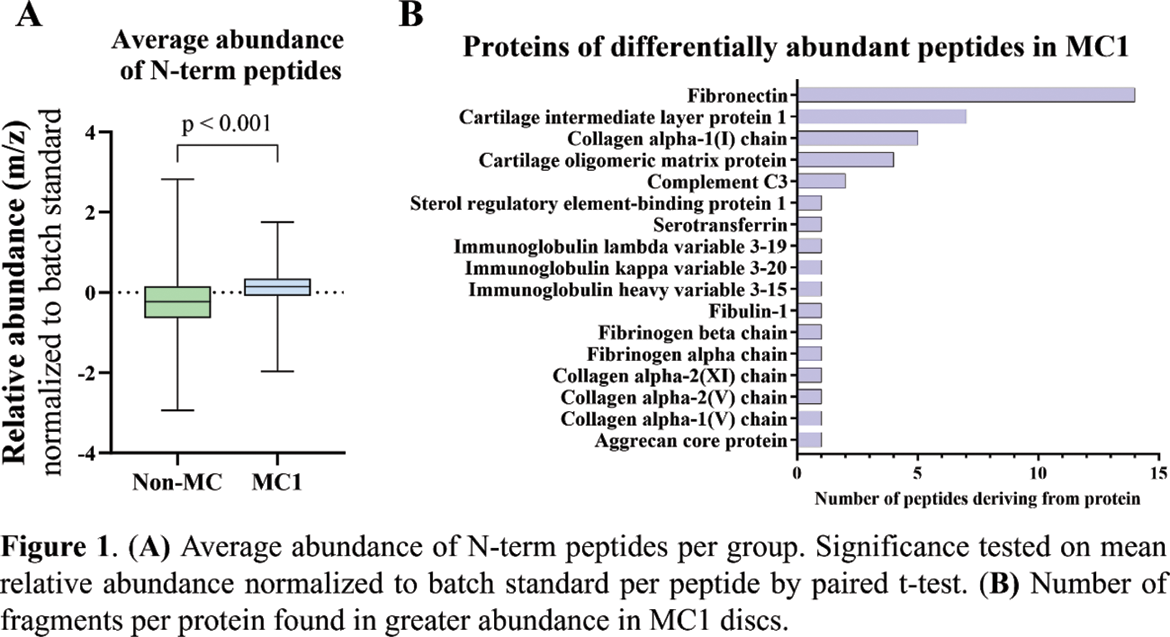
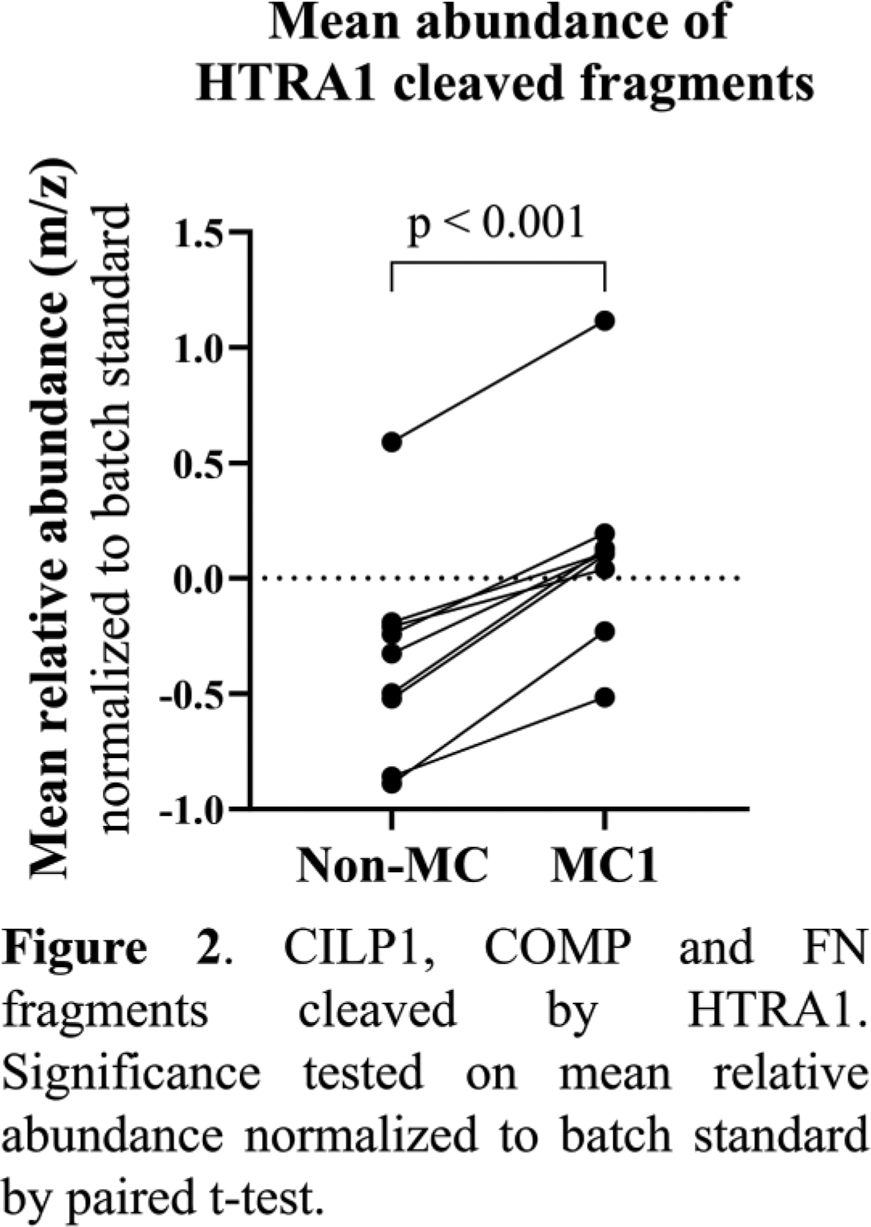
Results: The discs were assessed for degeneration using the Pfirrmann grade, revealing no significant difference between non-MC and MC1 discs (p = 0.370). Shotgun proteomics HTRA1 as the only protease with increased abundance in the MC1 discs. With N-TAILS, we found that overall, newly cleaved fragments were more abundant in MC1 discs, indicating a difference in degeneration patterns (Figure 1A). Among the 44 fragments found in greater abundance in MC1, 14 came from fibronectin (FN) cleavage, 7 from cartilage intermediate layer protein 1 (CILP1), 5 from type 1 collagen alpha-1 (COL1A1), and 4 from cartilage oligomeric matrix protein (COMP) (Figure 1B). In-vitro , HTRA1 cleaved recombinant CILP1, COMP and FN but not COL1A1. Analysis of the cleaved proteins with N-TAILS identified 20 N-terminally labeled peptides of which nine were also found in the ex vivo analyzed degenerated discs and were significantly more abundant in MC1 discs (Figure 2). FN fragments identified were part of the 29kDa N-terminal FN fragment, a known damage-associated molecular pattern.
Conclusion: We identified HTRA1 cleavage as a MC1-specific disc degradation mechanism contributing to increased ECM fragmentation in MC1 discs. Interestingly, HTRA1 is also highly relevant in osteoarthritis progression. The presence of ECM protein fragments has the potential to trigger pro-inflammatory and pro-fibrotic mechanisms. Given the higher concentration of these fragments in MC1 discs, it becomes crucial to investigate their possible role in the induction of inflammation associated with degenerative processes.
REFERENCES: NIL.
Acknowledgements: We would like to thank the FOREUM foundation for supporting our research.
Disclosure of Interests: Tamara Mengis: None declared, Bernd Roschitzki: None declared, Irina Heggli: None declared, Nick Herger: None declared, Mazda Farshad: None declared, Oliver Distler 4P-Pharma, Abbvie, Acceleron, Alcimed, Altavant, Amgen, AnaMar, Argenx, Arxx, AstraZeneca, Blade, Bayer, Boehringer Ingelheim, Corbus, CSL Behring, Galderma, Galapagos, Glenmark, Gossamer, Horizon, Janssen, Kymera, Lupin, Medscape, Merck, Miltenyi Biotec, Mitsubishi Tanabe, Novartis, Orion, Prometheus, Redxpharma, Roivant, Topadur and UCB., 4P-Pharma, Abbvie, Acceleron, Alcimed, Altavant, Amgen, AnaMar, Argenx, Arxx, AstraZeneca, Blade, Bayer, Boehringer Ingelheim, Corbus, CSL Behring, Galderma, Galapagos, Glenmark, Gossamer, Horizon, Janssen, Kymera, Lupin, Medscape, Merck, Miltenyi Biotec, Mitsubishi Tanabe, Novartis, Orion, Prometheus, Redxpharma, Roivant, Topadur and UCB., Stefan Dudli: None declared.