

Background GWAS have identified genetic variants associated with many complex diseases. These primarily affect regulatory elements that can affect expression of distal genes through chromatin interaction mechanisms. Previous studies have used functional genomics and chromatin conformation capture techniques to study the functional impact of these genetic variants [1]. However, such studies have predominantly used data generated from cell lines, which behave differently from primary cells. Additionally, there is a lack of research in Psoriatic Arthritis (PsA), which is a highly heterogenous disease that has been understudied and for which there are few effective treatments.
Objectives We aimed to study how chromatin conformation and gene expression is altered in primary T cells derived from patients with PsA, and to identify how genetic variants alter gene regulatory mechanisms in a population of individuals.
Methods Genotype data, together with matching high resolution Hi-C, RNA-seq and ATAC-seq was generated from primary CD4+ and CD8+ T cell samples isolated from the blood (n=55) and synovial fluid (n=9) of patients with PsA and blood of healthy controls (n=19).
Results We found that chromatin conformation is highly cell-type and context specific. Out of 105,956 total loops, 29,454 were differentially interacting (FDR<0.1) between CD8+ and CD4+ T cells, and 15,597 (FDR<0.1) between CD8+ T cells isolated from synovial fluid compared to those from blood. These differential loops overlap many disease associated GWAS loci such as the TNIP1 and STAT2 PsA loci. We identified 312 genotype associated loops, one of which is associated with a risk locus (GSDMB) known to be associated with several autoimmune diseases. Moreover, we found that chromatin conformation correlates strongly with differences in gene expression and chromatin accessibility, particularly with genes related to T cell activation and inflammatory pathways.
Next, we compared T cells isolated from blood of PsA patients to controls, but few significant differences were identified (Table 1). Segregating the PsA cohort by disease activity and treatment led to the identification of significantly larger differences. We found numerous differentially expressed genes (DEGs) and differential ATAC-seq peaks in patients with or without active disease, and in those receiving or not receiving biological disease-modifying antirheumatic drugs (bDMARD) (Table 1). Furthermore, there were numerous DEGs and differential ATAC-seq peaks in bDMARD responders (n=8) vs non-responders (n=6) (Figure 1, Table 1). DEGs were enriched for pathways involving immune function such as IL-2 and IL-15 signalling. Differential ATAC-seq peaks were enriched for binding sites for transcription factors such as RUNX1, JUN, IRF4 and STAT2. Furthermore, one of the differential ATAC-seq peaks we identified overlapped SNPs in the psoriasis RUNX3 locus.
| Comparison | RNA-seq CD4+ | RNA-seq CD8+ | ATAC-seq CD4+ | ATAC-seq CD8+ |
|---|---|---|---|---|
| Patients vs healthy controls | 46 | 200 | 0 | 4 |
| Patients with active disease vs remission | 169 | 36 | 1675 | 3 |
| bDMARD treatment effect in non-active disease | 90 | 13 | 75 | 14 |
| bDMARD treatment effect active disease | 967 | 295 | 13176 | 5069 |
| Remission vs active disease without bDMARD | 17 | 21 | 0 | 0 |
| Remission vs active disease treated with bDMARD | 1034 | 374 | 2559 | 41 |
Image/graph:
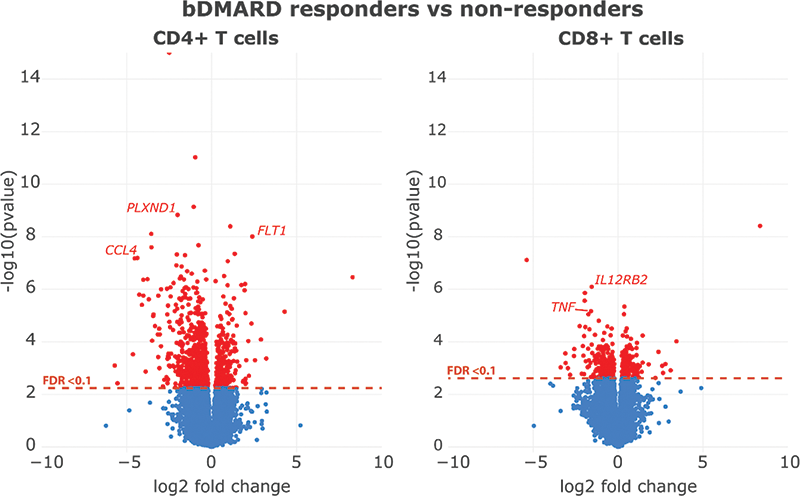
Figure 1. Volcano plot of DEGs between bDMARD responders vs non-responders in CD4+ and CD8+ T cells respectively.
Conclusion This study provides novel insight into gene regulatory mechanisms, particularly regarding PsA GWAS signals and disease pathogenesis. We identified, for the first time, epigenetic differences correlated with disease severity and treatment response in PsA using primary T cells.
Reference [1]Orozco, Gisela et al. “3D genome organization links non-coding disease-associated variants to genes.” Frontiers in cell and developmental biology vol. 10 995388. 20 Oct. 2022
Acknowledgements The authors would like to acknowledge the assistance given by IT Services and the use of the Computational Shared Facility at The University of Manchester.
This work was funded by the Wellcome Trust (award references 207491/Z/17/Z and 215207/Z/19/Z), Versus Arthritis (award reference 21754), NIHR Manchester Biomedical Research Centre and the Medical Research Council (award reference MR/N00017X/1).
Disclosure of Interests None Declared.
Keywords: Psoriatic arthritis, Genetics/Epigenetics, -omics
DOI: 10.1136/annrheumdis-2023-eular.981