

Background The enthesis is a the key target tissue in spondyloarthritis (SpA) with inflammation evident in both the enthesis soft tissue (EST) and the adjacent anchoring peri-entheseal bone (PEB)[1]. The healthy human spinal enthesis harbour, resident mesenchymal stem cells (MSCs), myeloid cells, innate immune cell populations including γδ T cells, innate lymphoid cells (ILCs), conventional T-cells but a lack of FOXP3+ regulatory T cells (Treg) [2, 3]. Given the predominance of stroma in EST and the known immunomodulatory effects of MSCs it is credible that such cells could assume important enthesis location immunomodulatory function.
Objectives To investigate the single cell RNAseq (scRNAseq) landscape of EST and PEB tissues. Reflecting RNAseq findings we examined immunomodulatory capacity of MSCs derived from EST and PEB in co-cultures with allogeneic CD3/CD28-stimulated CD4+ T cells.
Methods For scRNAseq, interspinous ligament and spinous process samples from patients undergoing spinal surgery (n=4). Samples were divided into EST and PEB and enzymatically digested. Following viability enrichment, we performed single cell partitioning with 10x Chromium, followed by oligo-barcoded RNA library generation and Illumina sequencing. For the co-culture, EST and PEB were expanded until passage 2 (n=5) using bone marrow MSCs (BM-MSCs) as control standard (n=2). CD4+ CD25- T-cells from blood were induced to proliferate using CD3/CD28 stimulation with MSCs/CD4+/CD25- cell co-cultured at 1:1, 2:1, 4:1, and 8:1 ratios for 5 days. T cells proliferation index was calculated and Treg related markers expressed on MSCs including CD73 and CD39 were measured in PEB and EST.
Results Following quality control, the transcriptome of 25,635 single cells was analysed. The compositional analysis revealed different abundance, such as stromal cells being more abundant in the EST, and haematopoietic precursors almost exclusively in the PEB. Analysis of the main cell populations revealed: the presence of innate and innate-like lymphocyte subpopulations such as MAIT and γδT cells, and a transcriptional gradient in neutrophils, reflecting various stages of maturation. In the T cell compartment, a putative regulatory (Tr1-like) population characterised by TGFB1 expression, was more abundant than conventional Foxp3+ Tregs. A number of stromal cells expressed the regulatory marker NT5E (CD73).). Consistently with scRNAseq, EST-MSCs expressed CD73 significantly more than PEB-MSCs prior to co-culture (p<0.001). with significant upregulation with T- cells coculture (p<0.05).
EST and PEB derived MSCs showed trilineage differentiation to bone, cartilage and fat[3] and significantly supressed CD4+ CD25- proliferation in dose dependent manner compared to control co-culture (p<0.001). Interestingly, PEB-MSCs showed an increase in the CD39 expression after co-culture with T cells compare to PEB-MSCs with no T cells at ratio 1:4 and 1:8 (P<0.001). No EST-MSCs CD39 expresion increase after T cells co-culture was noted but PEB-MSCs CD39 expression increased at 1:4 and 1:8 ratio (P<0.01, P<0.01 respectively.
Conclusion Our single-cell transcriptomic analysis suggests that EST and PEB have few Tregs and that other T cells and MSCs possess immunomodulatory functions in these tissues. This immunomodulatory activity may be important at these sites of high mechanical stress to maintain homeostasis.
References
Image/graph:
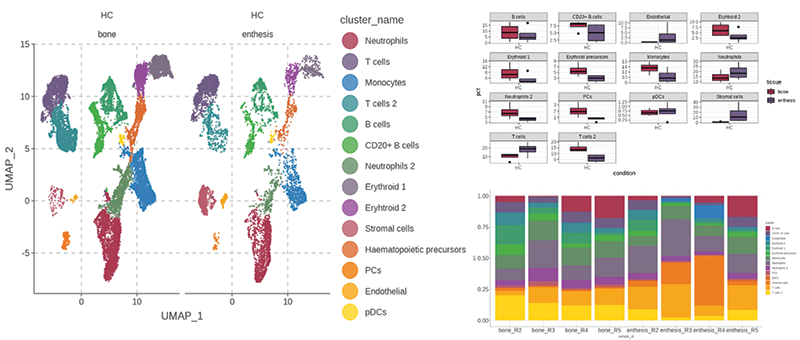
Figure 1. Healthy spinal enthesis dataset paired PEB and EST.A: cluster composition and anatomical distribution. B: Cluster abundance by tissue. C: Per sample cluster abundance
Acknowledgements: NIL.
Disclosure of Interests None Declared.
Keywords: Enthesitis, Inflammatory arthritides, Innate immunity
DOI: 10.1136/annrheumdis-2023-eular.5837