

Background Abatacept (ABA or CTLA4Ig) is a fusion protein that interferes with co-stimulation of T cells and antigen-presenting cells by inhibiting interactions between CD28 and CD80/86. ABA is effective in rheumatoid arthritis, but clinical trials of patients with Systemic Lupus Erythematosus (SLE) have produced disappointing results. Exploratory analyses of clinical subsets has led to the hypothesis that ABA might benefit some SLE patients depending on disease severity or outcome defnitions[1,2], but identification of reproducible predictors of response remains elusive.
Objectives We used data and samples collected from previous clinical trials to identify pathways of SLE pathogenesis and potential biomarkers which predict clinical response to ABA.
Methods Samples were obtained from 20 patients from the ABC study which evaluated ABA in SLE patients without organ-threatening disease on limited background medications and 46 patients from the ACCESS trial of ABA in active nephritis patients on a background of low dose cyclophosphamide followed by azathioprine. Neither trial met their key endpoints vs placebo. Pre-treatment gene expression profiles were compared in those who did or did not achieve clinical response to ABA. HSIC-Lasso[3] was employed to select the top 20 independent features from whole blood transcriptomic profiles which best distinguished responders and non-responders across both trials. B cells, monocytes, and T cells were isolated from ABC patient PBMCs for further cell type-specific transcriptomic profiling. Differentially expressed genes (DEGs) were identified in each cell type by DESeq2 at false discovery rate < 0.1.
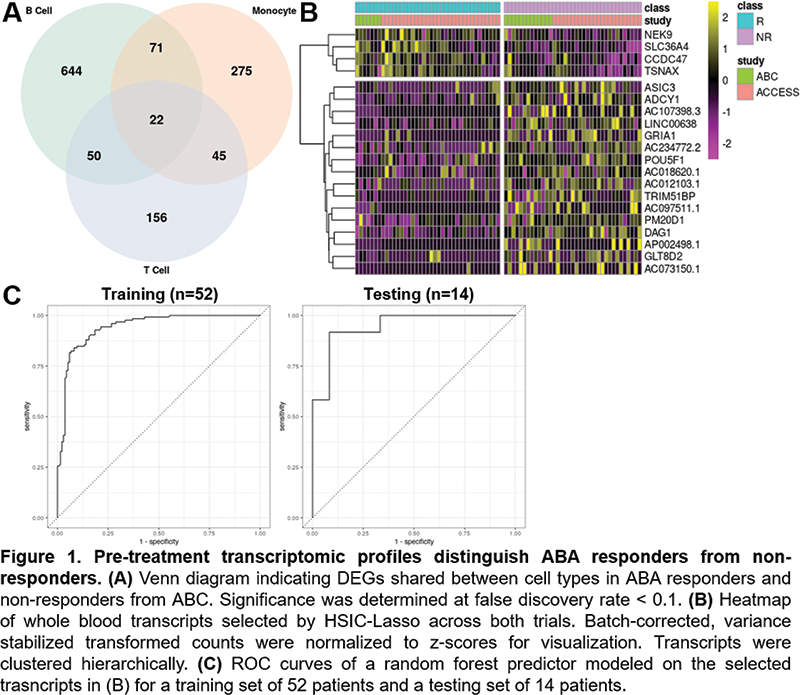
Results Pre-treatment DEGs associated with abatacept responders were cell type-specific, with the most distinctions observed in B cells (Figure 1). These DEGs included FCRLA, CXCR3, and TNFRSF17, indicating baseline differences in B cell activation and differentiation in responding patients. Baseline whole blood expression patterns could distinguish later response vs non-response to ABA in both clinical trials (Figure 1). Transcripts identified by HSIC-Lasso included multiple unannotated lncRNAs and pseudogenes, as well as transcription factors, transporters, and enzymatic components of cell structure and metabolism. Random forest modeling and receiver operating characteristics (ROC) were employed to test the ability of differentiating transcripts to predict response (Figure 1). 5-fold cross validation of the predictor on a training set of 52 samples had 85.8 (± 2.2) % accuracy with ROC AUC: 0.939 (± 0.0166). 14 confirmation samples were similarly modeled and achieved an accuracy of 83.3% with ROC AUC: 0.944.
Conclusion The pathophysiology of SLE is heterogeneous and complex, but approaches to predict treatment response by patterns of gene expression may help identify patients most likely to benefit from a given treatment.
References
Image/graph:
Acknowledgements Samples and data from the ACCESS trial were provided by the Immune Tolerance Network.
Disclosure of Interests Kevin Thomas: None declared, Miles Smith: None declared, Carla Guthridge: None declared, Nicolas Dominguez: None declared, Susan Macwana: None declared, Wade DeJager: None declared, Peter Schafer: None declared, Stan Kamp: None declared, Aikaterini Thanou: None declared, Cristina Arriens: None declared, Joan Merrill Consultant of: AbbVie, Alexion, Alumis, Amgen, Astra Zeneca, Aurinia, Bristol Myers Squibb, EMD Serono, Genentech, Gilead, GlaxoSmithKline, Lilly, Merck, Pfizer, Provention, Remegen, Sanofi, UCB, and Zenas., Grant/research support from: Astra Zeneca, Bristol Myers Squibb, and GlaxoSmithKline., Judith A. James: None declared, Joel Guthridge: None declared.
Keywords: Clinical Trials, Systemic lupus erythematosus, Biomarkers
DOI: 10.1136/annrheumdis-2023-eular.2064