

Background Systemic lupus erythematosus (SLE) is the prototypical systemic autoimmune rheumatic disease and is characterized by hyperactive B cells leading to autoantibody production. Changes in gut microbiota are associated with autoimmune disease including SLE. We theorize that changes in the gut microbiota seen in patients with SLE relate to both serological, clinical manifestations as well as disease activity in SLE. A better understanding of the gut microbiota may lead to its use as a biomarker, part of disease activity scoring in SLE, or a potential future therapeutic target.
Objectives The objective was to determine the relationship between changes in the gut microbiota and the patient reported outcomes using patient global health assessment (PG) and overall disease assessment using Systemic Lupus Erythematosus Disease Activity Index 2000 (SLEDI). Secondary objective including the relationship between serological markers of SLE, and proteinuria was also investigated.
Methods A total of 36 patients were enrolled, and 20 patients who met the Systemic Lupus Collaborating Clinics (SLICC) criteria for SLE each provided a stool sample. Table 1 demonstrates some of the patient and disease related information collected. The stool samples were homogenized, and the DNA was extracted. The V4 region of 16S rRNA gene was amplified using PCR. The purified amplicons were sequenced bi-directionally. Samples were analyzed using the R package phyloseq. Reads were processed and amplicon sequence variants (ASVs) were generated using DADA2 in R. Taxonomy was assigned using the SILVA 138 rRNA database training set using the DADA2 functions, assignTaxonomy and addSpecies. Two samples with fewer than 1000 reads were removed from the dataset, resulting in 18 samples and 1311 ASVs used for downstream microbiota and statistical analyses.
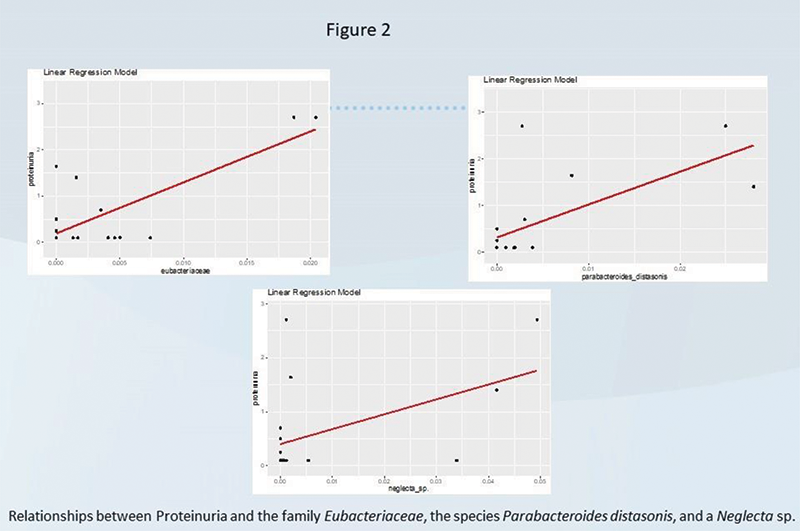
Results There was also no statistical relationship between the gut microbiota and patient global health assessment or SLEDI. We found no statistical correlation between the gut microbiota and lymphopenia, complement C3/C4, ANA titers, SSA/SSB results or the presence of lupus nephritis. There was an inverse relationship between the family of Ruminococcaceae and dsDNA antibody levels (Figure 2). We also noted significant positive correlation between the level of proteinuria and the genus Eubacterium (family Eubacteriaceae) and the species Parabacteroides distasonis and a Neglecta sp. (Figure 2).
Conclusion There were significant changes in gut microbiota in SLE that correlated with the levels of dsDNA antibodies and proteinuria. The findings are suggestive that there may be a direct relationship between lupus nephritis serology, clinical symptoms, and the gut microbiota.
| Age | Race | Sex | dsDNA | C3/C4 | Proteinuria | SLEDAI-2K | PGA | LN |
|---|---|---|---|---|---|---|---|---|
| 18 | W | F | No | No | <0.1 | 0 | 50 | No |
| 41 | W | M | Elevated | No | <0.1 | 2 | 0 | Yes |
| 21 | A | F | Elevated | Low | 1.64 | 8 | 0 | Yes |
| 66 | W | F | Elevated | No | <0.1 | 0 | 0 | No |
| 39 | AA | F | No | Low | 0.25 | 0 | 0 | No |
| 29 | AA | F | Elevated | Low | <0.1 | 2 | 15 | No |
| 65 | W | F | Elevated | Low | 0.7 | 13 | 50 | Yes |
| 27 | AA | M | Elevated | Low | 0.5 | 6 | 10 | Yes |
| 36 | W | M | No | No | <0.1 | 0 | 5 | No |
| 25 | W | F | Elevated | Low | 1.4 | 8 | 15 | Yes |
| 64 | AA | F | No | Low | <0.1 | 6 | 10 | No |
| 23 | W | M | Elevated | No | <0.1 | 2 | 0 | Yes |
| 56 | W | F | No | No | <0.1 | 1 | 0 | No |
| 40 | AA | F | No | No | 2.7 | 4 | 20 | Yes |
| 35 | AA | F | No | No | <0.1 | 0 | 0 | No |
| 37 | AA | F | Elevated | No | <0.1 | 0 | 0 | No |
| 39 | AA | F | No | No | 2.7 | 5 | 50 | Yes |
| 47 | W | F | No | No | <0.1 | 0 | 0 | No |
| 53 | W | F | Elevated | Low | 0.1 | 0 | 0 | Yes |
dsDNA = Double Stranded DNA antibody, C3 = Complement, C4 = Complement 4, SLEDAI-2K = Systemic Lupus Erythematosus Disease Activity Index 2000, PGA = Physician Global assessment, LN = Lupus nephritis, Race: W=White, A= Asia, AA, African American, Sex: F= Female, M= Male
Image/graph:
References
Acknowledgements: NIL.
Disclosure of Interests None Declared.
Keywords: Autoantibodies, Systemic lupus erythematosus, Kidneys
DOI: 10.1136/annrheumdis-2023-eular.278