

Background: Psoriatic Arthritis (PsA) is a heterogeneous inflammatory disease of which the exact pathobiology remains insufficiently understood. Proteomics or other -omics can be a powerful tool to investigate biological processes or to endotype patients, but interpretation of results can be complicated by the influence of many patient and disease characteristics, including treatment used.
Objectives: The aim of the current analysis was to identify 1) differences in proteomic profile between newly diagnosed, disease modifying antirheumatic drugs (DMARD) naïve, PsA patients and those previously failing DMARD-therapy (DMARD-failure) 2) potential mediation of these differences by patient, disease and treatment characteristics 3) biological pathways that are differentially expressed between PsA subtypes.
Methods: Baseline data from the TOFA-PREDICT trial discovery cohort (40 DMARD-naïve and 40 DMARD-failure PsA patients) were used [1]. Proteomic profiling of 368 proteins (cardiovascular III, cardiometabolic, inflammation and metabolism panels) was performed on serum using Olink Proteomics technology. Differences were studied using univariate Welch 2-sample t-test or Mann-Whitney U with FDR corrected p-values and multivariate models (sPLS-DA and XGBoost). Mediation analysis was done to test if differences in proteins can be explained by age, sex, BMI, arthritis/skin/enthesitis markers or MTX/corticosteroid use. Enrichment analysis of markers was conducted (Gene Ontology function annotation, Kyoto Encyclopedia of Genes and Genomes pathway analysis, and DisGeNET disease enrichment) and a protein-protein interaction (PPI) network was constructed using StringDB [2,3].
Results: Expression of 16 proteins was significantly downregulated in DMARD-failures (DAB2, NRP1, LILRB1, TIMP1, TNXB, PTPRS, GP1BA, METRNL, MFAP5, VASN, CA3, IGFBP6, DPP4, SERPINA7, VCAM1, CD59) and 3 were upregulated (CD5, SCGB3A2, LTBP2). Multivariate approaches also identified these 19 proteins, next to 15 other key markers that differentiate DMARD-failure and DMARD-naïve patients (IL-1RT2, PGLYRP1, CCL4, ST6GAL1, ANXA11, CDH5, CD8A, KIT, ADGRE2, ITGB7, Gal-3, MMP-10, GDF-15, PLXNB2, Ep-CAM).
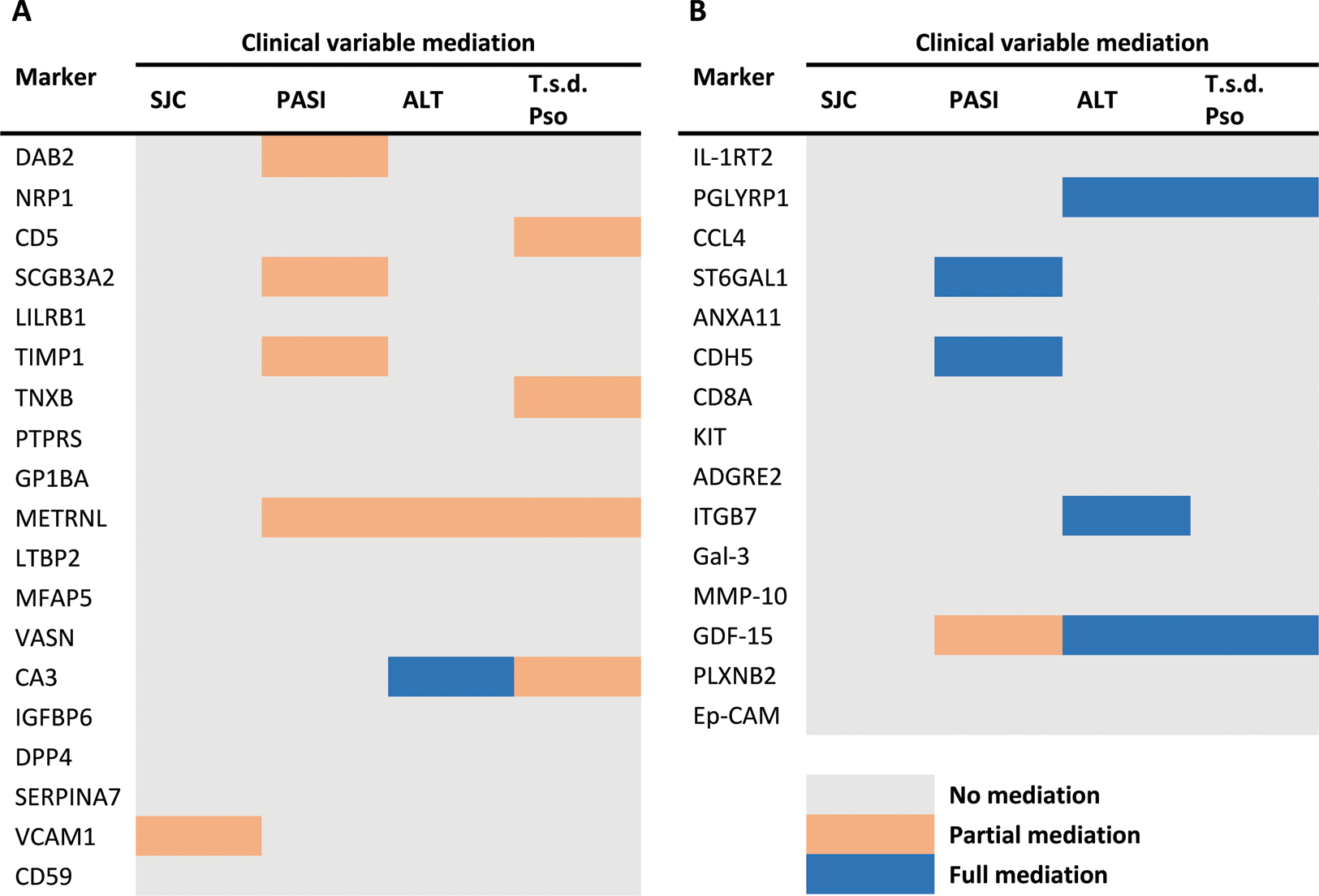
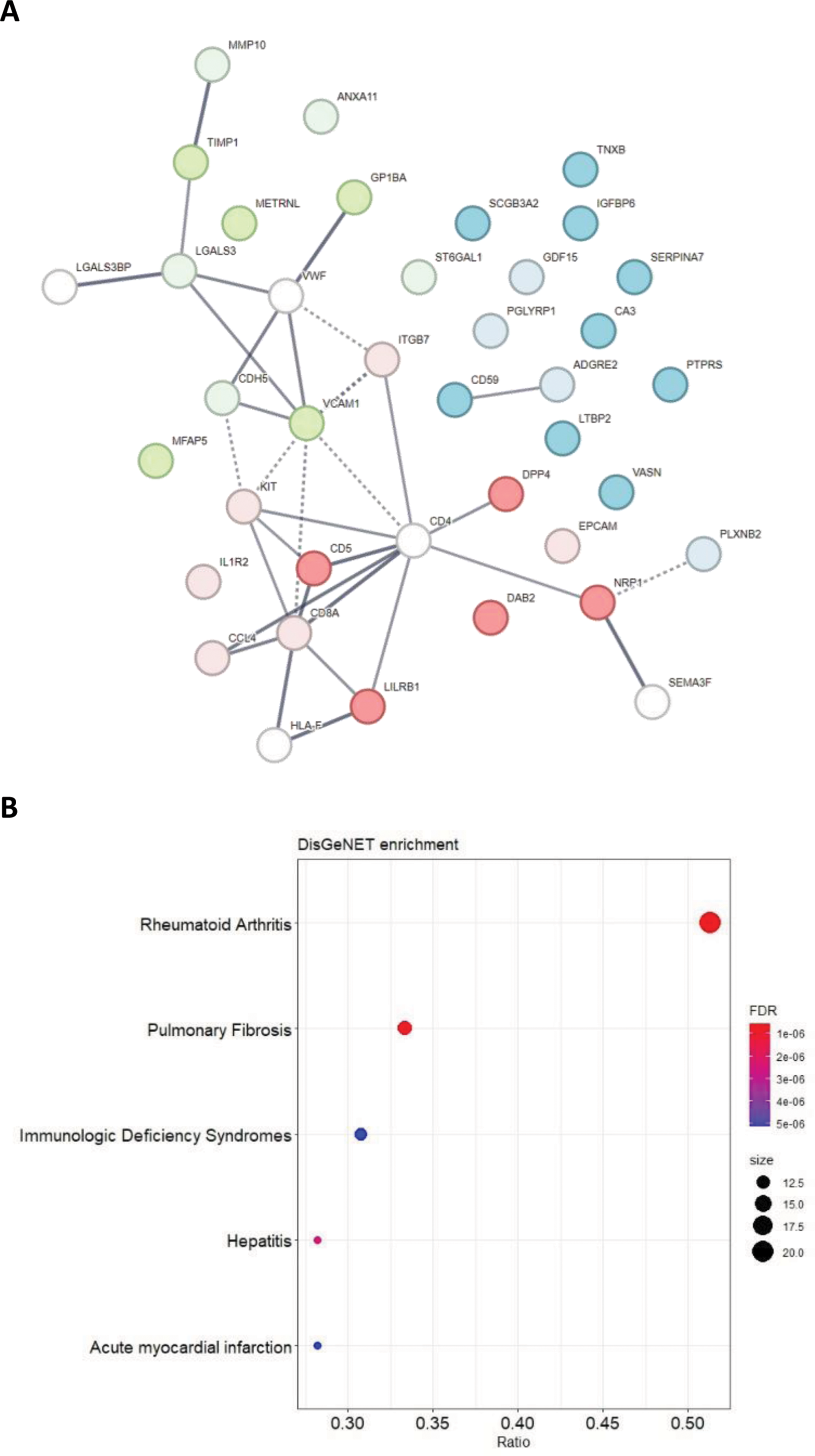
Mediation analysis indicated that clinical characteristics or MTX/corticosteroid use did not fully mediate the difference between the two patient groups, except for 6 markers (CA3, PGLYRP1, ST6GAL1, CDH5, ITGB7, GDF-15) that were mediated by either Psoriasis Area and Severity Index, ALT level or time since diagnosis of psoriasis (Table 1). The PPI network and enrichment analysis showed that proteins are mainly involved in hematopoietic cell lineage and cell adhesion molecules pathways and are associated with some pathologies, such as rheumatoid arthritis, autoimmune disease and hematopoietic system disease (Figure 1).
Conclusion: This study demonstrates proteomic differences between PsA patients that are DMARD-naïve and those previously failing DMARD therapy. This should be considered in proteomic analysis involving both type of patients. Only some of these differences were (partly) mediated, mostly by extend/duration of skin involvement and liver damage marker (ALT). Thus, using proteomic profiles, clinical classification of PsA patients and understanding of pathophysiological mechanisms underlying PsA could be improved. We are currently evaluating metabolic profiles with a similar approach to further extend this research.
REFERENCES: [1] Kleinrensink, Nienke J., et al. “TOFA-PREDICT study protocol: a stratification trial to determine key immunological factors predicting tofacitinib efficacy and drug-free remission in psoriatic arthritis (PsA).” BMJ open 12.10 (2022): e064338.
[2] Szklarczyk, Damian, et al. “STRING v10: protein–protein interaction networks, integrated over the tree of life.” Nucleic acids research 43.D1 (2015): D447-D452.
[3] Piñero, Janet, et al. “The DisGeNET knowledge platform for disease genomics: 2019 update.” Nucleic acids research 48.D1 (2020): D845-D855.
Acknowledgements: This study was conducted as a collaboration between University Medical Center Utrecht and Pfizer. Pfizer is the study sponsor and the project is co-funded by the PPP Allowance made available by Health~Holland Top Sector Life Sciences & Health, to stimulate public–private partnerships (grant number: LSHM17074).
Disclosure of Interests: Mieke Bentvelzen: None declared, Said el Bouhaddani: None declared, Simon C. Mastbergen: None declared, Julia Spierings Research grants from Boehringer Ingelheim and Miltenyi Biotec., Harald E. Vonkeman Paid consultant for (pharmaceutical) companies Abbvie, Novartis, Pfizer, UCB, Johnson and Johnson, Financial grants from: Galapagos and Boehringer Ingelheim, Shasti C. Mooij: None declared, Lydia G. Schipper: None declared, Amin Herman: None declared, Floris Lafeber: None declared, Hae-Won Uh: None declared, Paco M.J. Welsing: None declared